Dendritic spines (Yuste)
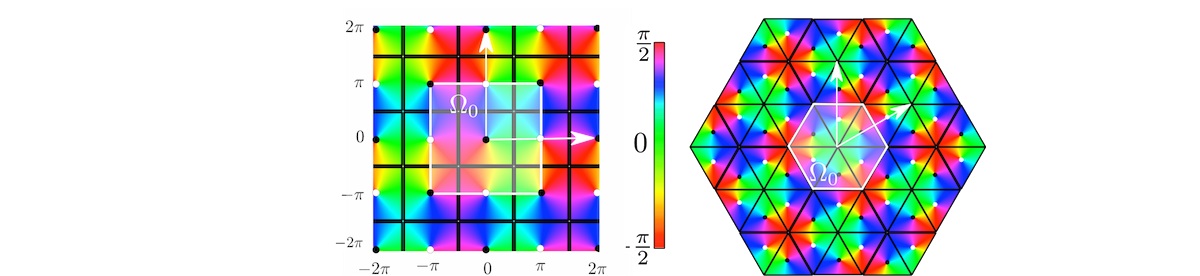
PO map
Brainbow (Tamily Weissmann)
Contact me for the following programs. Alternatively, you may look at public repositories on bitbucket, github, gitlab accounts.
Bifurcation Analysis code in Julia. Allows to perform continuation and (automatic) bifurcation analysis in large dimensions on CPUs / GPUs. ODEs, DDEs, PDEs are also well within the scope of the package. The organisation bifurcationkit regroups many plugins, for example to follow homoclinic orbits.
Simulation of PDMPs in Julia based on this paper with speed on par with C. PDMP means piecewise deterministic Markov process.
A Julia wrapper to call the LSODA algorithm by Linda Petzold and Alan Hindmarsh. It solves ODE by switching automatically between stiff and non-stiff methods.
Simulation code (asic_model_container.pdf) in Neuron associated with our paper posted on BioRxiv. Download the file (right click, Save Link As...), change the name into asic_model_container.zip and unzip it.
I have been working with Bill Spotz from the Sandia Labs to develop PyTrilinos, e.g. the nonlinear / numerical continuation part, in Trilinos > v.12.
Simulation of Neural Fields equations using petsc4py. This allows parallel and fast simulations of models $$\frac{d}{dt}V(x,t) = -V(x,t) + \int_\Omega J(x,y)S(V(y,t))dy,\ y\in\mathbb R^2$$
together with the numerical bifurcation analysis. The number of unknowns can be >$10^6$.
Computing Hopf bifurcation curves for delay differential equations (paper), basically finding $\lambda\in i\mathbb R$ such that there is a solution in $U$ to
$$ \left(\lambda+l\right) U_i(x)=\sum\limits_{j=1}^p\int\limits_\Omega J_{ij}(x,y)e^{-\lambda\tau(x,y)}U_j(y)dy,\ 1\leq i\leq p,\ x,y\in\mathbb R^d$$